 |
Chaste
Release::2017.1
|
 |
Chaste
Release::2017.1
|
#include <AbstractBackwardEulerCardiacCell.hpp>
 Inheritance diagram for AbstractBackwardEulerCardiacCell< SIZE >:
Inheritance diagram for AbstractBackwardEulerCardiacCell< SIZE >: Collaboration diagram for AbstractBackwardEulerCardiacCell< SIZE >:
Collaboration diagram for AbstractBackwardEulerCardiacCell< SIZE >:Public Member Functions | |
| AbstractBackwardEulerCardiacCell (unsigned numberOfStateVariables, unsigned voltageIndex, boost::shared_ptr< AbstractStimulusFunction > pIntracellularStimulus) | |
| virtual | ~AbstractBackwardEulerCardiacCell () |
| virtual void | ComputeResidual (double time, const double rCurrentGuess[SIZE], double rResidual[SIZE])=0 |
| virtual void | ComputeJacobian (double time, const double rCurrentGuess[SIZE], double rJacobian[SIZE][SIZE])=0 |
| OdeSolution | Compute (double tStart, double tEnd, double tSamp=0.0) |
| void | ComputeExceptVoltage (double tStart, double tEnd) |
| void | SolveAndUpdateState (double tStart, double tEnd) |
 Public Member Functions inherited from AbstractCardiacCell Public Member Functions inherited from AbstractCardiacCell | |
| AbstractCardiacCell (boost::shared_ptr< AbstractIvpOdeSolver > pOdeSolver, unsigned numberOfStateVariables, unsigned voltageIndex, boost::shared_ptr< AbstractStimulusFunction > pIntracellularStimulus) | |
| virtual | ~AbstractCardiacCell () |
| void | Init () |
| void | SetTimestep (double dt) |
| void | SetVoltage (double voltage) |
| double | GetVoltage () |
| unsigned | GetNumberOfStateVariables () const |
| unsigned | GetNumberOfParameters () const |
| std::vector< double > | GetStdVecStateVariables () |
| const std::vector< std::string > & | rGetStateVariableNames () const |
| void | SetStateVariables (const std::vector< double > &rVariables) |
| void | SetStateVariable (unsigned index, double newValue) |
| void | SetStateVariable (const std::string &rName, double newValue) |
| double | GetAnyVariable (const std::string &rName, double time=0.0) |
| double | GetParameter (const std::string &rParameterName) |
| double | GetParameter (unsigned parameterIndex) |
| void | SetParameter (const std::string &rParameterName, double value) |
| virtual void | SetState (CellModelState state) |
| virtual void | SetSlowValues (const std::vector< double > &rSlowValues) |
| virtual void | GetSlowValues (std::vector< double > &rSlowValues) |
| virtual bool | IsFastOnly () |
| virtual void | AdjustOutOfRangeSlowValues (std::vector< double > &rSlowValues) |
| virtual unsigned | GetNumSlowValues () |
 Public Member Functions inherited from AbstractCardiacCellInterface Public Member Functions inherited from AbstractCardiacCellInterface | |
| AbstractCardiacCellInterface (boost::shared_ptr< AbstractIvpOdeSolver > pOdeSolver, unsigned voltageIndex, boost::shared_ptr< AbstractStimulusFunction > pIntracellularStimulus) | |
| virtual | ~AbstractCardiacCellInterface () |
| virtual double | GetIIonic (const std::vector< double > *pStateVariables=NULL)=0 |
| unsigned | GetVoltageIndex () |
| void | SetStimulusFunction (boost::shared_ptr< AbstractStimulusFunction > pStimulus) |
| double | GetStimulus (double time) |
| void | SetIntracellularStimulusFunction (boost::shared_ptr< AbstractStimulusFunction > pStimulus) |
| double | GetIntracellularStimulus (double time) |
| double | GetIntracellularAreaStimulus (double time) |
| void | SetUsedInTissueSimulation (bool tissue=true) |
| virtual boost::shared_ptr< RegularStimulus > | UseCellMLDefaultStimulus () |
| bool | HasCellMLDefaultStimulus () |
| virtual AbstractLookupTableCollection * | GetLookupTableCollection () |
| boost::shared_ptr< AbstractStimulusFunction > | GetStimulusFunction () |
| const boost::shared_ptr< AbstractStimulusFunction > | GetStimulusFunction () const |
| const boost::shared_ptr< AbstractIvpOdeSolver > | GetSolver () const |
| void | SetSolver (boost::shared_ptr< AbstractIvpOdeSolver > pSolver) |
| virtual void | SetVoltageDerivativeToZero (bool clamp=true) |
| void | SetFixedVoltage (double voltage) |
| virtual void | SetStretch (double stretch) |
| virtual double | GetIntracellularCalciumConcentration () |
 Public Member Functions inherited from AbstractOdeSystem Public Member Functions inherited from AbstractOdeSystem | |
| AbstractOdeSystem (unsigned numberOfStateVariables) | |
| virtual | ~AbstractOdeSystem () |
| virtual bool | CalculateStoppingEvent (double time, const std::vector< double > &rY) |
| virtual double | CalculateRootFunction (double time, const std::vector< double > &rY) |
| bool | GetUseAnalyticJacobian () |
| const std::vector< double > & | rGetConstStateVariables () const |
 Public Member Functions inherited from AbstractParameterisedSystem< std::vector< double > > Public Member Functions inherited from AbstractParameterisedSystem< std::vector< double > > | |
| AbstractParameterisedSystem (unsigned numberOfStateVariables) | |
| std::vector< double > & | rGetStateVariables () |
| std::vector< double > | GetStateVariables () |
| void | SetStateVariables (const std::vector< double > &rStateVariables) |
| double | GetStateVariable (unsigned index) const |
| double | GetStateVariable (const std::string &rName) const |
| void | SetStateVariable (unsigned index, double newValue) |
| void | SetStateVariable (const std::string &rName, double newValue) |
| virtual void | VerifyStateVariables () |
| void | SetDefaultInitialConditions (const std::vector< double > &rInitialConditions) |
| void | SetDefaultInitialCondition (unsigned index, double initialCondition) |
| std::vector< double > | GetInitialConditions () const |
| void | ResetToInitialConditions () |
| double | GetParameter (unsigned index) const |
| double | GetParameter (const std::string &rName) const |
| void | SetParameter (const std::string &rName, double value) |
| void | SetParameter (unsigned index, double value) |
| double | GetAnyVariable (unsigned index, double time=0.0, std::vector< double > *pDerivedQuantities=NULL) |
| double | GetAnyVariable (const std::string &rName, double time=0.0, std::vector< double > *pDerivedQuantities=NULL) |
| void | SetAnyVariable (unsigned index, double value) |
| void | SetAnyVariable (const std::string &rName, double value) |
| virtual std::vector< double > | ComputeDerivedQuantities (double time, const std::vector< double > &rState) |
| std::vector< double > | ComputeDerivedQuantitiesFromCurrentState (double time) |
 Public Member Functions inherited from AbstractUntemplatedParameterisedSystem Public Member Functions inherited from AbstractUntemplatedParameterisedSystem | |
| AbstractUntemplatedParameterisedSystem (unsigned numberOfStateVariables) | |
| virtual | ~AbstractUntemplatedParameterisedSystem () |
| boost::shared_ptr< const AbstractOdeSystemInformation > | GetSystemInformation () const |
| std::string | GetSystemName () const |
| unsigned | GetNumberOfAttributes () const |
| bool | HasAttribute (const std::string &rName) const |
| double | GetAttribute (const std::string &rName) const |
| unsigned | GetNumberOfStateVariables () const |
| const std::vector< std::string > & | rGetStateVariableNames () const |
| const std::vector< std::string > & | rGetStateVariableUnits () const |
| unsigned | GetStateVariableIndex (const std::string &rName) const |
| bool | HasStateVariable (const std::string &rName) const |
| std::string | GetStateVariableUnits (unsigned index) const |
| unsigned | GetNumberOfParameters () const |
| const std::vector< std::string > & | rGetParameterNames () const |
| const std::vector< std::string > & | rGetParameterUnits () const |
| unsigned | GetParameterIndex (const std::string &rName) const |
| bool | HasParameter (const std::string &rName) const |
| std::string | GetParameterUnits (unsigned index) const |
| unsigned | GetNumberOfDerivedQuantities () const |
| const std::vector< std::string > & | rGetDerivedQuantityNames () const |
| const std::vector< std::string > & | rGetDerivedQuantityUnits () const |
| unsigned | GetDerivedQuantityIndex (const std::string &rName) const |
| bool | HasDerivedQuantity (const std::string &rName) const |
| std::string | GetDerivedQuantityUnits (unsigned index) const |
| unsigned | GetAnyVariableIndex (const std::string &rName) const |
| bool | HasAnyVariable (const std::string &rName) const |
| std::string | GetAnyVariableUnits (unsigned index) const |
| std::string | GetAnyVariableUnits (const std::string &rName) const |
Protected Member Functions | |
| virtual void | ComputeOneStepExceptVoltage (double tStart)=0 |
| virtual void | UpdateTransmembranePotential (double time)=0 |
 Protected Member Functions inherited from AbstractParameterisedSystem< std::vector< double > > Protected Member Functions inherited from AbstractParameterisedSystem< std::vector< double > > | |
| std::string | DumpState (const std::string &rMessage) |
| std::string | DumpState (const std::string &rMessage, std::vector< double >Y) |
| std::string | DumpState (const std::string &rMessage, std::vector< double >Y, double time) |
| void | CheckParametersOnLoad (const std::vector< double > &rParameters, const std::vector< std::string > &rParameterNames) |
Private Member Functions | |
| template<class Archive > | |
| void | serialize (Archive &archive, const unsigned int version) |
| void | EvaluateYDerivatives (double time, const std::vector< double > &rY, std::vector< double > &rDY) |
Friends | |
| class | boost::serialization::access |
Additional Inherited Members | |
 Protected Attributes inherited from AbstractCardiacCell Protected Attributes inherited from AbstractCardiacCell | |
| double | mDt |
 Protected Attributes inherited from AbstractCardiacCellInterface Protected Attributes inherited from AbstractCardiacCellInterface | |
| unsigned | mVoltageIndex |
| boost::shared_ptr< AbstractIvpOdeSolver > | mpOdeSolver |
| boost::shared_ptr< AbstractStimulusFunction > | mpIntracellularStimulus |
| bool | mSetVoltageDerivativeToZero |
| bool | mIsUsedInTissue |
| bool | mHasDefaultStimulusFromCellML |
| double | mFixedVoltage |
 Protected Attributes inherited from AbstractOdeSystem Protected Attributes inherited from AbstractOdeSystem | |
| bool | mUseAnalyticJacobian |
 Protected Attributes inherited from AbstractParameterisedSystem< std::vector< double > > Protected Attributes inherited from AbstractParameterisedSystem< std::vector< double > > | |
| std::vector< double > | mStateVariables |
| std::vector< double > | mParameters |
 Protected Attributes inherited from AbstractUntemplatedParameterisedSystem Protected Attributes inherited from AbstractUntemplatedParameterisedSystem | |
| unsigned | mNumberOfStateVariables |
| boost::shared_ptr< AbstractOdeSystemInformation > | mpSystemInfo |
This is the base class for cardiac cells solved using a (decoupled) backward Euler approach (see http://dx.doi.org/10.1109/TBME.2006.879425).
The basic approach to solving such models is:
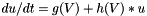 . The update expression is then
. The update expression is then 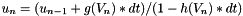 .
. 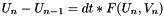 . The template parameter to the class specifies the size of this nonlinear system.
. The template parameter to the class specifies the size of this nonlinear system. Definition at line 67 of file AbstractBackwardEulerCardiacCell.hpp.
| AbstractBackwardEulerCardiacCell< SIZE >::AbstractBackwardEulerCardiacCell | ( | unsigned | numberOfStateVariables, |
| unsigned | voltageIndex, | ||
| boost::shared_ptr< AbstractStimulusFunction > | pIntracellularStimulus | ||
| ) |
Standard constructor for a cell.
| numberOfStateVariables | the size of the ODE system |
| voltageIndex | the index of the variable representing the transmembrane potential within the state variable vector |
| pIntracellularStimulus | the intracellular stimulus function |
Some notes for future reference:
Definition at line 208 of file AbstractBackwardEulerCardiacCell.hpp.
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::serialize().
|
virtual |
Virtual destructor
Definition at line 219 of file AbstractBackwardEulerCardiacCell.hpp.
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::serialize().
|
virtual |
Simulates this cell's behaviour between the time interval [tStart, tEnd], with timestep mDt. Uses a forward Euler step to update the transmembrane potential at each timestep.
The length of the time interval must be a multiple of the timestep.
| tStart | beginning of the time interval to simulate |
| tEnd | end of the time interval to simulate |
| tSamp | sampling interval for returned results (defaults to mDt) |
Reimplemented from AbstractCardiacCell.
Definition at line 223 of file AbstractBackwardEulerCardiacCell.hpp.
References AbstractBackwardEulerCardiacCell< SIZE >::ComputeOneStepExceptVoltage(), AbstractCardiacCell::mDt, AbstractUntemplatedParameterisedSystem::mpSystemInfo, OdeSolution::rGetSolutions(), AbstractParameterisedSystem< std::vector< double > >::rGetStateVariables(), OdeSolution::rGetTimes(), OdeSolution::SetNumberOfTimeSteps(), OdeSolution::SetOdeSystemInformation(), AbstractBackwardEulerCardiacCell< SIZE >::UpdateTransmembranePotential(), and AbstractParameterisedSystem< std::vector< double > >::VerifyStateVariables().
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::serialize().
|
virtual |
Simulates this cell's behaviour between the time interval [tStart, tEnd], with timestep mDt. The transmembrane potential is kept fixed throughout.
The length of the time interval must be a multiple of the timestep.
| tStart | beginning of the time interval to simulate |
| tEnd | end of the time interval to simulate |
Reimplemented from AbstractCardiacCell.
Definition at line 275 of file AbstractBackwardEulerCardiacCell.hpp.
References AbstractBackwardEulerCardiacCell< SIZE >::ComputeOneStepExceptVoltage(), AbstractCardiacCell::mDt, and AbstractParameterisedSystem< std::vector< double > >::VerifyStateVariables().
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::serialize().
|
pure virtual |
Compute the Jacobian matrix for the nonlinear system portion of the cell model.
| time | the current time |
| rCurrentGuess | the current guess for 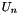 |
| rJacobian | to be filled in with the Jacobian matrix |
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::serialize().
|
protectedpure virtual |
Compute the values of all state variables, except the voltage, using backward Euler, for one timestep from tStart.
| tStart | start of this timestep |
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::Compute(), AbstractBackwardEulerCardiacCell< 0u >::Compute(), AbstractBackwardEulerCardiacCell< SIZE >::ComputeExceptVoltage(), AbstractBackwardEulerCardiacCell< 0u >::ComputeExceptVoltage(), AbstractBackwardEulerCardiacCell< SIZE >::EvaluateYDerivatives(), AbstractBackwardEulerCardiacCell< 0u >::EvaluateYDerivatives(), AbstractBackwardEulerCardiacCell< SIZE >::SolveAndUpdateState(), and AbstractBackwardEulerCardiacCell< 0u >::SolveAndUpdateState().
|
pure virtual |
Compute the residual of the nonlinear system portion of the cell model.
| time | the current time |
| rCurrentGuess | the current guess for 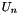 |
| rResidual | to be filled in with the residual vector |
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::serialize().
|
inlineprivatevirtual |
This function should never be called - the cell class incorporates its own solver.
| time | |
| rY | |
| rDY |
Implements AbstractOdeSystem.
Definition at line 170 of file AbstractBackwardEulerCardiacCell.hpp.
References AbstractBackwardEulerCardiacCell< SIZE >::ComputeOneStepExceptVoltage(), NEVER_REACHED, and AbstractBackwardEulerCardiacCell< SIZE >::UpdateTransmembranePotential().
|
inlineprivate |
Archive the member variables.
| archive | |
| version |
Definition at line 79 of file AbstractBackwardEulerCardiacCell.hpp.
References AbstractBackwardEulerCardiacCell< SIZE >::AbstractBackwardEulerCardiacCell(), AbstractBackwardEulerCardiacCell< SIZE >::Compute(), AbstractBackwardEulerCardiacCell< SIZE >::ComputeExceptVoltage(), AbstractBackwardEulerCardiacCell< SIZE >::ComputeJacobian(), AbstractBackwardEulerCardiacCell< SIZE >::ComputeResidual(), AbstractBackwardEulerCardiacCell< SIZE >::SolveAndUpdateState(), and AbstractBackwardEulerCardiacCell< SIZE >::~AbstractBackwardEulerCardiacCell().
|
virtual |
Simulate this cell's behaviour between the time interval [tStart, tEnd], with timestemp mDt, updating the internal state variable values.
| tStart | beginning of the time interval to simulate |
| tEnd | end of the time interval to simulate |
Reimplemented from AbstractCardiacCell.
Definition at line 301 of file AbstractBackwardEulerCardiacCell.hpp.
References TimeStepper::AdvanceOneTimeStep(), AbstractBackwardEulerCardiacCell< SIZE >::ComputeOneStepExceptVoltage(), TimeStepper::GetTime(), TimeStepper::IsTimeAtEnd(), AbstractCardiacCell::mDt, AbstractBackwardEulerCardiacCell< SIZE >::UpdateTransmembranePotential(), and AbstractParameterisedSystem< std::vector< double > >::VerifyStateVariables().
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::serialize().
|
protectedpure virtual |
Perform a forward Euler step to update the transmembrane potential.
| time | start of this timestep |
Referenced by AbstractBackwardEulerCardiacCell< SIZE >::Compute(), AbstractBackwardEulerCardiacCell< 0u >::Compute(), AbstractBackwardEulerCardiacCell< SIZE >::EvaluateYDerivatives(), AbstractBackwardEulerCardiacCell< 0u >::EvaluateYDerivatives(), AbstractBackwardEulerCardiacCell< SIZE >::SolveAndUpdateState(), and AbstractBackwardEulerCardiacCell< 0u >::SolveAndUpdateState().
|
friend |
Needed for serialization.
Definition at line 71 of file AbstractBackwardEulerCardiacCell.hpp.